Note
Click here to download the full example code
Plotting Neural Network Training Curves¶
This is a basic example using a convolutional recurrent neural network to learn segments directly from time series data
# Author: David Burns
# License: BSD
import matplotlib.pyplot as plt
import numpy as np
from tensorflow.python.keras.layers import Dense, LSTM, Conv1D
from tensorflow.python.keras.models import Sequential
from tensorflow.python.keras.wrappers.scikit_learn import KerasClassifier
from pandas import DataFrame
from sklearn.model_selection import train_test_split
from seglearn.datasets import load_watch
from seglearn.pipe import Pype
from seglearn.transform import Segment
Simple NN Model¶
def crnn_model(width=100, n_vars=6, n_classes=7, conv_kernel_size=5,
conv_filters=3, lstm_units=3):
input_shape = (width, n_vars)
model = Sequential()
model.add(Conv1D(filters=conv_filters, kernel_size=conv_kernel_size,
padding='valid', activation='relu', input_shape=input_shape))
model.add(LSTM(units=lstm_units, dropout=0.1, recurrent_dropout=0.1))
model.add(Dense(n_classes, activation="softmax"))
model.compile(loss='categorical_crossentropy', optimizer='adam',
metrics=['accuracy'])
return model
Setup¶
# load the data
data = load_watch()
X = data['X']
y = data['y']
# split the data
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.25, random_state=42)
# create a segment learning pipeline
pipe = Pype([('seg', Segment(width=100, step=100, order='C')),
('crnn', KerasClassifier(build_fn=crnn_model, epochs=4, batch_size=256,
verbose=0, validation_split=0.2))])
Accessing training history¶
# this is a bit of a hack, because history object is returned by the
# keras wrapper when fit is called
# this approach won't work with a more complex estimator pipeline, in which case
# a callable class with the desired properties should be made passed to build_fn
pipe.fit(X_train, y_train)
history = pipe.history.history
print(DataFrame(history))
# depends on version
if 'accuracy' in history:
ac_train = history['accuracy']
ac_val = history['val_accuracy']
elif 'acc' in history:
ac_train = history['acc']
ac_val = history['val_acc']
else:
raise ValueError("History object doesn't contain accuracy record")
epoch = np.arange(len(ac_train)) + 1
Out:
loss accuracy val_loss val_accuracy
0 1.957218 0.184432 1.948542 0.047619
1 1.950467 0.192847 1.946699 0.036415
2 1.943282 0.189341 1.944778 0.028011
3 1.937308 0.205470 1.943990 0.022409
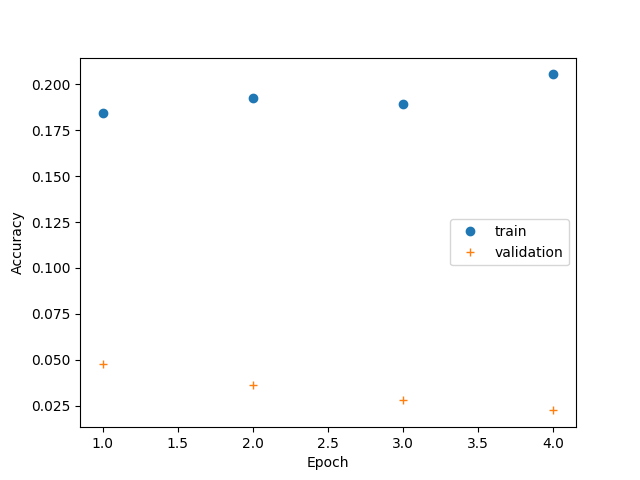
Training Curves¶
plt.plot(epoch, ac_train, 'o', label="train")
plt.plot(epoch, ac_val, '+', label="validation")
plt.xlabel("Epoch")
plt.ylabel("Accuracy")
plt.legend()
plt.show()
Out:
/home/david/Code/seglearn/examples/plot_nn_training_curves.py:96: UserWarning: Matplotlib is currently using agg, which is a non-GUI backend, so cannot show the figure.
plt.show()
Total running time of the script: ( 0 minutes 3.716 seconds)